We are at siege. A siege by an unknown enemy. An enemy with which we are befuddled. And unless you were living under a rock for the past couple of months(like Jared Leto), you know what I’m talking about – COVID-19. Whether you turn on the news, or scroll through social media, the majority of information that you take in nowadays is about the SARS-COV2 virus, or the Novel Corona Virus.
But among all the negativity, there was a sliver of light shining through. When faced with a common enemy, mankind united across borders(for the most part; there are bad apples always) to help each other tide over the current assault. Scientists, who are the heroes of the day, doubled down to find a cure, vaccine, and a million other things which helps in the battle against COVID-19. And along with the real heroes, Data Scientists were also called to action to help in any way they could. A lot of people tried their best at forecasting the progression of the disease, so that the Governments can plan better. A lot more dedicated their time in analysing the data coming out of a multitude of sources to prepare dashboards, or network diagrams, etc. to help understand the progression of the disease. And yet another set of people tried to apply the techniques of AI to things like identifying the risk of a patient, or help diagnose the disease with X-Rays, etc.
While following these developments, one particular area where a lot of people did some attempt is Chest Radiography based identification of COVID-19 cases. One of those early attempts received a lot of attention, volunteers, funding etc. along with a lot of flak for the positioning the research took(You can read more about it here). TLDR; A PhD candidate out of Australia used a pretrained model(Resnet50), trained on 50 images, messed up the code because of train validation leak, and claimed 97% accuracy on COVID-19 case identification. Some others even got a 100% accuracy(turned out it was trained on the same dataset on which it got a 100% accuracy).
Along with this noise, there was an arxiv preprint came out of University of Waterloo, Canada by Linda Wang and Alexander Wong titled, COVID-Net: A Tailored Deep Convolutional Neural Network Design for Detection of COVID-19 Cases from Chest Radiography Images. In the paper, they propose a new CNN Architecture which was trained from scratch on a dataset of 5941 posteroanterior chest radiography images. To generate the dataset, they combined two publicly available datasets – COVID chest X-ray dataset, and Kaggle Chest X-ray images (pneumonia) dataset. In the paper, they divided this dataset into four classes – COVID-19, Viral, Bacterial, and Normal. The below bar chart shows the class distribution of train and test splits. This was a decently sized dataset, although the COVID cases were on the lower side. They reported a 100% Recall and an 80% Precision for the model.
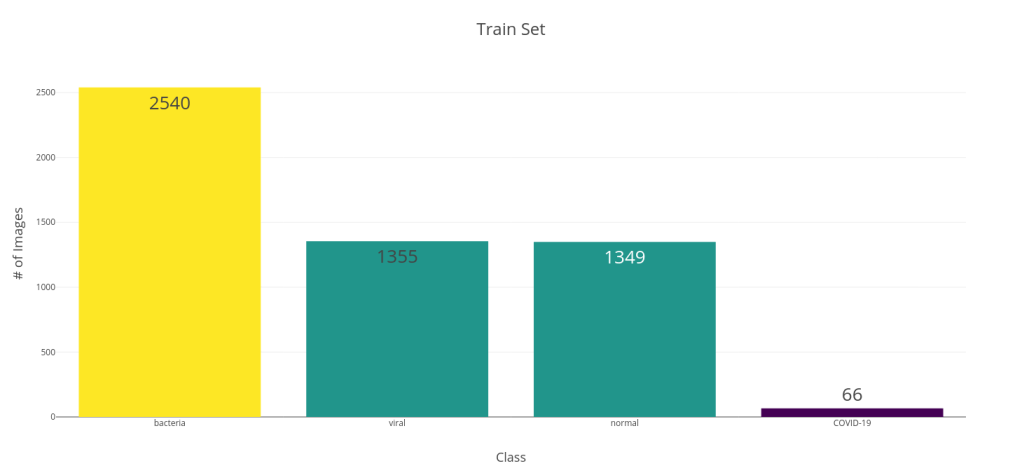

This was the first dataset of decent size on COVIDx and it got me interested. And since they shared the trained model and code to evaluate in a Github Repo, this was prime for an analysis.
Disclaimer
I feel I need to state a disclaimer right about here. Whatever follows is a purely academic exercise and not at all an attempt to suggest this as a verifiable and valid way of testing for COVID-19. First things first. I personally do not endorse this attempt at identifying COVID-19 using any of these models. I very little knowledge about medicine, and absolutely zero about reading an X-ray. Unless this has been verified and vetted by a medical professional, this is nothing better than a model trained on a competition data set.
There are a few problems also regarding the dataset.
- COVID-19 cases and the other cases comes from different data sources and it’s doubtful if the model is identifying the data source or actual visual indicators representing COVID-19. I’ve tried to look at GradCAM results, but me being an absolute zero in reading an X-ray, I don’t know if the model is looking at the right indicators.
- It is also unclear as to what stage a patient was when the X-ray was taken. If it was something that was taken too late in his disease, this method does not hold it’s ground.
Why not Transfer Learning?
The first thought I had when I saw the model and the dataset was this – Why not Transfer Learning? The dataset is quite small, especially the class that we are interested in. Training a model from scratch and trying to properly capture the complex representation of the different classes, especially the COVID-19 class was a little bit of a stretch for me.
But playing the Devil’s advocate, why would a CNN trained on animals and food (the most popular classes in ImageNet) do better in X-rays? The network trained on natural and colourful images might have learnt totally different feature representations necessary than what is necessary to handle monochromatic X-rays.
As a rational human being and a staunch believer of the process of Science, I decided to look up the existing literature on it. Surprisingly, the research community is divided about the issue. Let me make it clear. There is no debate as to whether pretraining or Transfer Learning works for medical images. But the debate is about whether pretraining on Imagenet has any benefit. There were papers which claimed Imagenet pretraining helped Medical Image Classification and segmentation. And there were papers who pushed for random initialisation for the weights.
There was a recent paper by Veronika Cheplygina[1] which did a review of the literature in the perspective of whether or not Imagenet pretraining is beneficial for medical images. The conclusion was – “It depends”. Another recent paper from Google Brain(which was accepted into NeurIPS 2019)[2] deep dives into this issue. Although the general conclusion was that transfer learning with Imagenet weights is not beneficial for medical imaging, they do point out a few other interesting behaviour:
- When the sample size is small, as is most cases in medical imaging, Transfer Learning, even if it is Imagenet based, is beneficial
- When they look at the convergence rates of the different models, pretrained ones converged faster
- Most interesting result was that they tried initializing the networks with random weights, but derived the mean and standard deviation of the random initialization based on pretrained weights and found that it too provided the convergence speedup that pretrained models had.
Bottom line was that the study didn’t show worse performance for Imagenet trained models and had faster convergence. Even though the large Imagenet models may be over-parametrized for the problem, it does offer some benefit if you want to get a model working as fast as possible.
The Experiment
Now that I’ve done the literature review, it was time to test out my hypothesis. I gathered the dataset, wrote up a training script, and tested out a few Pretrained models.
| Model | # of Parameters | GFLOPS |
| DenseNet 121 | 8,010,116 | ~3 |
| Xception | 22,915,884 | – |
| ResNeXt 101 32x4d | 44,237,636 | ~7.8 |
I’ve used the FastAI library(a wrapper around PyTorch), which is very easy to use, especially if you are doing Transfer Learning with it’s easy “freeze” and “unfreeze” functions. Most of the experiments were run either on my laptop with a meagre GTX 1650 or on Google Colab. I’ve used the amazing library pretrainedmodels by Cadene as a source of my pretrained models apart from torchvision.
Data Augumentation
As our training dataset is relatively small and because it has two different sources of X-rays, I’ve used a few transformations as data augmentation. It both increases the dataset samples as well as give better generalization capabilities to the model. The transforms used are:
- Horizontal Flip
- Rotate
- Zoom
- Brightness
- Warp
- Cutout
Training Procedure
Below are the basic steps I’ve used for the training of these models. Full code is published on GitHub.
- Import the models and create a learner from fastai. fastai has a few inbuilt mechanism to cut and split pretrained models so that we can use a custom head and apply discriminative learning rates easily. for the models in torchvision, the cut and split are predefined in fastai. But for models that are loaded from outside torchvision, we need to define those as well. ‘cut’ tells fastai where to make the separation between the feature extractor part of the CNN and the classifier part so that it can replace it with a custom head. “split” tells fastai how to split the different blocks on the CNN so that each block can have different learning rates.
- Split the Train into Train and Validation using a StratifiedShuffleSplit
- Kept the loss as a standard CrossEntropy
- Freeze the feature extractor part of the CNN and train the model. I used the One-Cycle Learning Rate Scheduler proposed by Leslie Smith[3]. It is heavily advocated by Jeremy Howard in his fastai courses and is implemented in the fastai library.
- After the learning saturates, unfreeze the rest of the model and finetune the model. Whether to use One-Cycle scheduler or not and whether to use differential learning rates or not, was decided empirically by looking at the loss curves.
Tricks used in Training
Mixup
Mixup[4] is a form of data augmentation where we generate new examples by weighted linear interpolation of two existing examples.
is between 0 and 1. In practice, it is sampled from a beta distribution which is parametrised by
. Typically,
is between 0.1 to 0.4 where the effect of mixup is not too much that it leads to underfitting.
Progressive Resizing
For DenseNet 121, I tried doing Progressive Resizing as well, just to see if it gets me better results. Progressive Resizing is when we start training the network with a small image size and then use the weights learned from the smaller size image and start training on a bigger size image and in stages we move to higher resolution image sizes. I tried it in three stages – 64×64, 128×128, and 224×224.
The Results
Without further ado, let’s take a look at the results.
Densenet 121
Best DenseNet model was got by progressively resizing 64×64 –> 128×128 and using mixup during training.

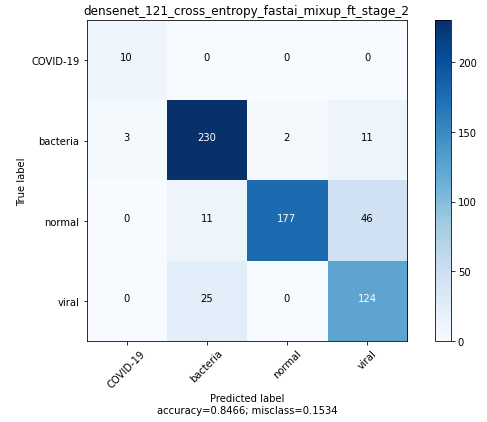
Xception
The best Xception model was trained using mixup and finetuned after initial pretraining with frozen weights.
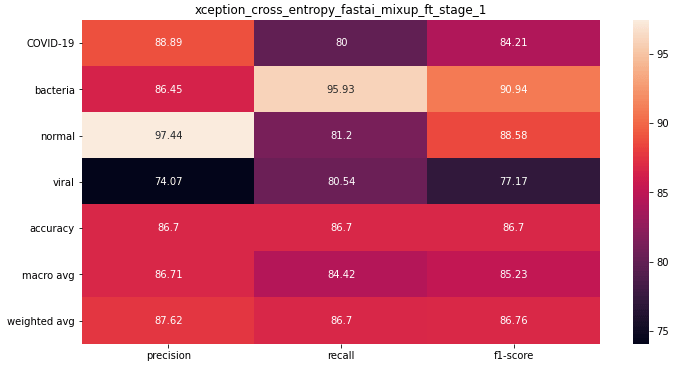
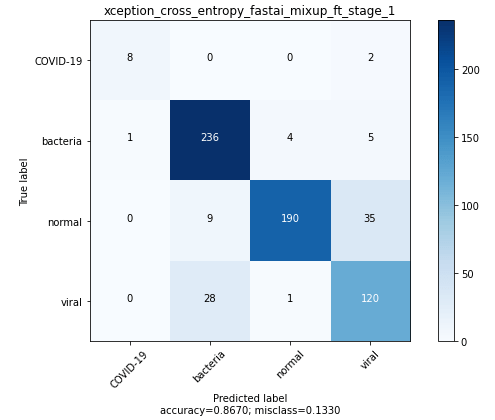
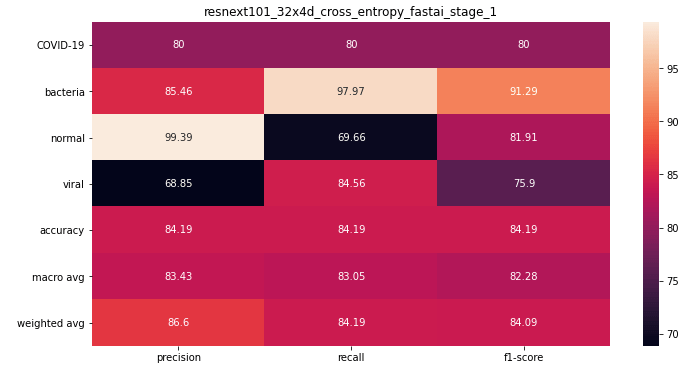
ResneXt 101 32x4d
The best ResNeXt model was trained without mixup(did not try), and without finetuning(finetuning was giving me worse performance for some reason).

Let’s summarize these results in a table and place them alongside the results from the COVID-Net paper.
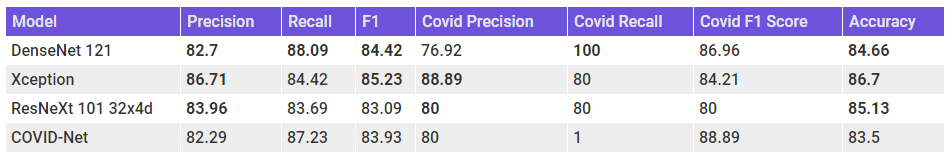
We can see right off the bat that all the models have a better accuracy than COVID-Net. But Accuracy isn’t the right metric to evaluate here. The Xception model, with the highest F1 score, seems to be the best performing model among the lot. But, if we look at Precision and Recall separately, we can see that COVID-Net is having high recall, especially for the COVID-19 cases, whereas our models have high Precision. Densenet 121 have a perfect recall, but the Precision is bad. But the Xception model has high precision and a not too bad recall.
Ensembling
We have seen that DenseNet was a high recall model and Xception was a high precision model. Would the performance be better if we average the predictions across both these models?

Not much different as before. Our ensemble still doesn’t have better recall. Let’s try a Weighted Ensemble to give more weight to Densenet which has a perfect recall in COVID-19. To determine the optimum weight, I use the predictions in the validation set and tried different weights.

Let’s add these ensembles also to the earlier table for comparison.
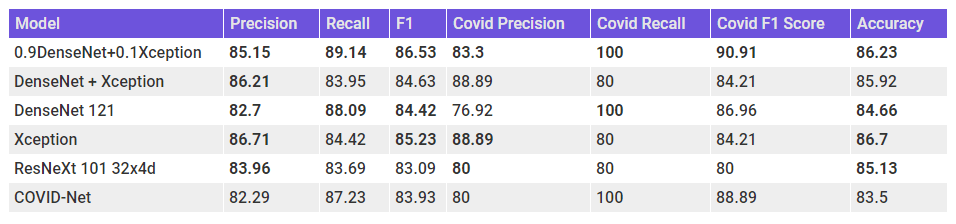
Finally, we have a model which has a balance between Precision and Recall and also beats the COVID-Net scores across all the metrics. Let’s take a look at the Confusion Matrix for the ensemble.
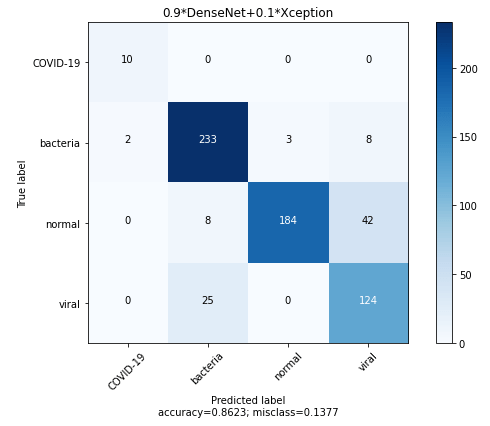
MODEL COMPLEXITY and Inference Time
When we are thinking about the usability of the model, we should also keep in mind the model complexity and inference time. The below table shows the number of parameters as a proxy for model complexity and inference time on my machine(GTX 1650).
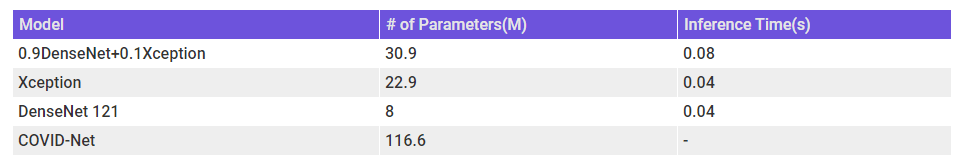
N.B. – Was not able to run inference on COVID-Net on my laptop(which has a terrible relationship with Tensorflow) and therefore do not know the inference time for the model. But by udging from the # of parameters, it should be more than the other models.
N.B. – The # of parameters and Inference time for the ensemble is taken as the summation of the constituents.
Model Analysis
GRAD CAM
Class Activation maps were introduced as a way to understand what a CNN is looking at while paking predictions way back in 2015 by Zhou, Bolei et al[5]. It is a technique to understand the regions of the image a CNN focuses on while making predictions. They achieve this by projecting back the weights of the output layer back to the output from the Convolutional Neural Network part.
Grad CAM is a generalization of CAM for many end use cases, apart from classification, and they acheive this by using the gradients w.r.t. the class at the last output from the Convolutional Layers. The authors of the paper[6] say:
Gradient-weighted Class Activation Mapping (Grad-CAM), uses the gradients of any target concept (say logits for ‘dog’ or even a caption), flowing into the final convolutional layer to produce a coarse localization map highlighting the important regions in the image for predicting the concept.
Let’s see a few examples of our predictions and their activations overlayed as a heatmap. Although I don’t understand if the network is looking at the right places, if somebody who is reading this know how to, reach out to me and let me know.
COVID-19
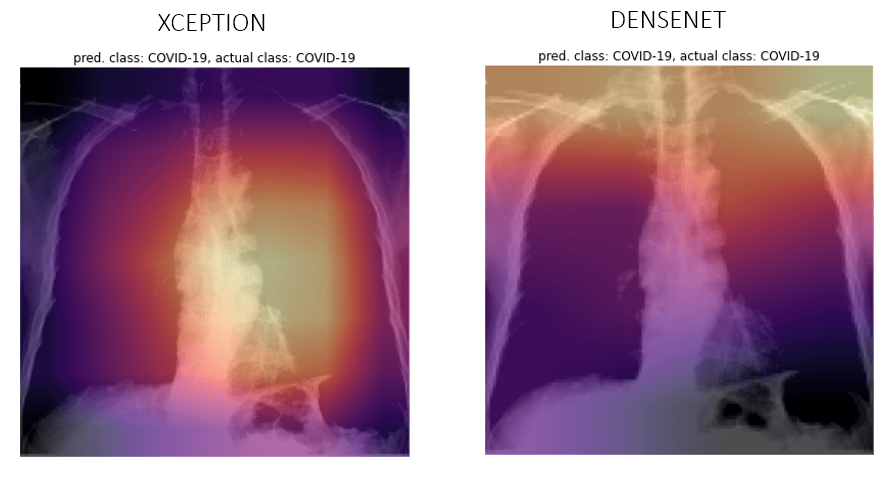
Viral
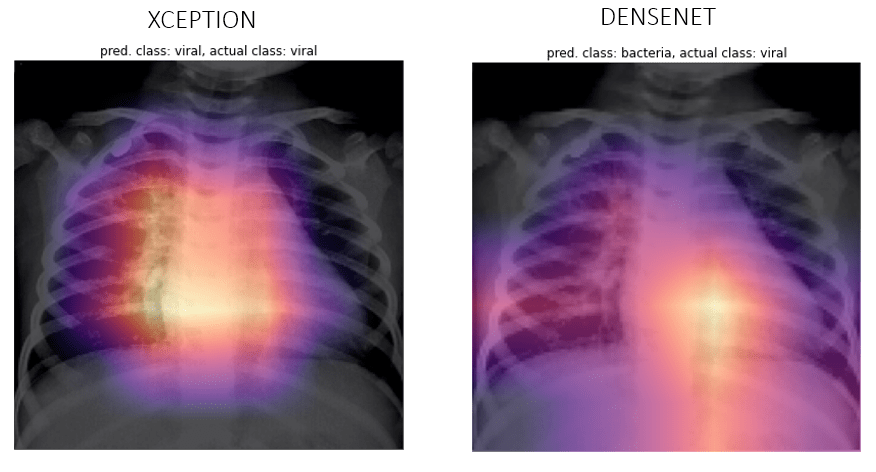
Bacteria

Normal

When Neural Networks Cheat

Feature Representation(t-SNE)
We can also take a look at how good the feature representations that come out of these networks are. Since the output from the Convolutional Layers are high dimensional, we’ll have to use a dimensionality reduction technique to plot it in two dimensions.
A popular method for exploring high-dimensional data is t-SNE, introduced by van der Maaten and Hinton in 2008[7]. t-SNE, unlike something like a PCA, isn;t a linear projection.It uses the local relationships between points to create a low-dimensional mapping. This allows it to capture non-linear structure.
Let’s take a look at the t-SNE vizualizations, with Perplexity 50 for the three models – COVID-Net, Xception, DenseNet.
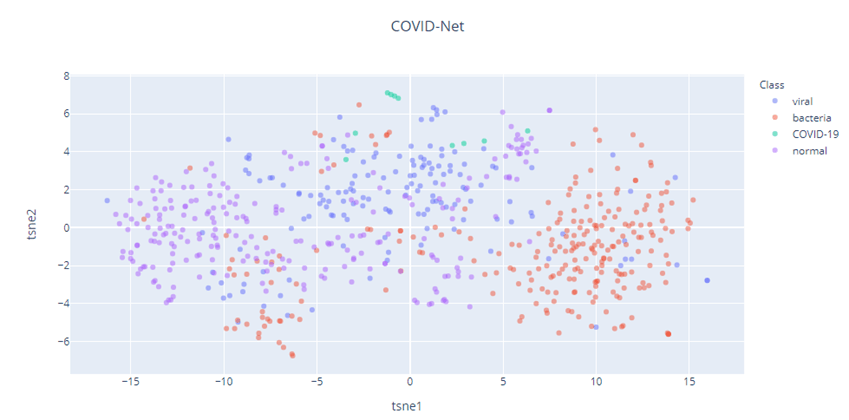
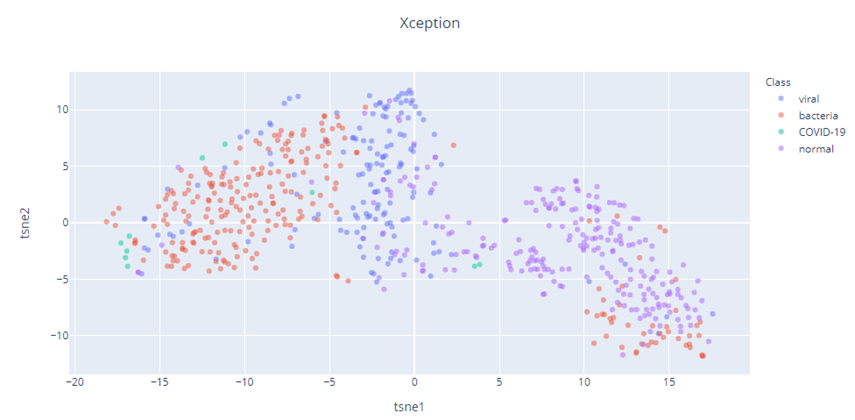

It also appears that our Imagenet pretrained models (Xception and DenseNet), has a better feature representation than COVID-Net. The t-SNE of COVID-Net is quite spread out and there is a lot of interspersion between the different classes. But the Xception and DenseNet feature representations show much better degree of separation of the different classes. The COVID-19 cases(Green) in all three cases shows separation, but because the dataset is so small, we need to take that inference with a grain of salt.
Final Words
We’ve seen that the Imagenet pretrained models performed better than the COVID-Net model. The best Xception model had better Precision and best DenseNet model had better Recall. In this particular scenario, Recall is what matters more because you need to be safe than sorry. Even if you classify a few non COVID-19 cases as positive, they will just be directed to a proper medical test. But the other kind of error is not that forgiving. So going purely by that, our DenseNet model is the better. But we also need to keep in mind that this has been trained on a limited data set. And that too, the number of COVID-19 cases were just around 60. It is highly likely that the model has memorised or overfit to those 60. A prime example of the case where the model used the label on the Xray to classify that as COVID-19. The GradCAM examination was also not very helpful, as some of the examples seemed like the model is looking at the right places. But for some examples, the heat map lit up almost all of the X-ray.
But after examining the GradCAM and the t-SNE, I think that the Xception model has learned a much better representation for the cases. The problem of having low Recall is something that can be dealt with.
On the larger point, with which we have started this whole exercise with, I think we can safely say that Imagenet pretraining does help in classification of Chest Radiography images of COVID-19(I did try to train DenseNet without pretrained weights on the same dataset without much sucess).
Unexplored Directions
There are a lot of unexplored dimensions to this problem and I’m gonna mention those in case any of my readers want to take those up.
- Collecting more data, especially COVID-19 cases, and retraining the models
- Dealing with the Class Imbalance
- Pretraining on Grayscale ImageNet[9] and subsequent Transfer Learning on Grayscale X-Rays
- Using the CheXpert dataset[8] as a bridge between Imagenet and Chest Radiography images by fine-tuning Imagenet models on CheXpert dataset and then apply to the problem at hand
Call for Collaboration
If you are a medical professional, who think this is a worthwhile direction of research, do reach out to me. I want to be convinced that this is effective, but currently am not.
If you are a ML researcher, who want to collaborate on publishing a paper, or continue this line of research, do reach out to me.
Update – After I’ve cloned the COVID-Net repo, there has been an update, both in the data set by adding a few more cases for COVID-19, and the addition of a larger model. And the performance of our ensemble is still better than the large model.
References
- Cheplygina, Veronika , “Cats or CAT scans: transfer learning from natural or medical image source datasets?,” arXiv:1810.05444 [cs.CV], Jan. 2019.
- Raghu, Maithra et.al, “Transfusion: Understanding Transfer Learning for Medical Imaging”, arXiv:1902.07208 [cs.CV], Feb. 2019
- Smith, Leslie N., “Cyclical Learning Rates for Training Neural Networks”, arXiv:1506.01186 [cs.CV], June.2019
- Hongyi Zhang, Moustapha Cissé, Yann N. Dauphin, David Lopez-Paz, “mixup: Beyond Empirical Risk Minimization”. ICLR (Poster) 2018
- Zhou, Bolei et al, “Learning Deep Features for Discriminative Localization”, arXiv:1512.04150, Dec. 2015
- Selvaraju, Ramaprasaath R. et al, “Grad-CAM: Visual Explanations from Deep Networks via Gradient-based Localization”. arXiv:1610.02391 [cs.CV], Oct. 2016
- Laurens van der Maaten, Geoffrey Hinton, “Visualizing Data using t-SNE”. 2008
- Irvin, Jeremy et al. “CheXpert: A Large Chest Radiograph Dataset with Uncertainty Labels and Expert Comparison”, arXiv:1901.07031 [cs.CV], Jan, 2019
- Xie, Yiting & Richmond, David. (2019). Pre-training on Grayscale ImageNet Improves Medical Image Classification: Munich, Germany, September 8-14, 2018, Proceedings, Part VI. 10.1007/978-3-030-11024-6_37.